12 Sep 2016
Chronic Wasting Disease (CWD), also in Norway
For those of you who read Norwegian see here for synopsis of the brief (at the time of writing it is 12-09-2016), yet dramatic, history of CWD in Norway.
In short, this spring one reindeer was discovered with CWD during capture for the reindeer monitoring project in Nordfjella. A few weeks later, two moose were found with CWD in another area further North.
This triggered a large surveying effort (underway) of deer in Norway together with the hunters this fall (starting with the reindeer hunt on September 20th). In addition, also the 318 reindeer killed by lightening in Hardangervidda were screened for CWD (all negative). The first results are both positive and negative. It seems that CWD does not occur in the reindeer population at high incidence. Unfortunately, one reindeer shot in the Nordfjella area was found to have CWD, indicating that the one in spring wasn’t an isolated case.
Thus, to date four animals (2 reindeer & 2 moose) have been found with CWD in Norway. The moose hunt is still to start, so in a couple months we will have a much better idea of the full extent of CWD in Norway. It seems, however, that this disease will be a factor for population management in the years to come. This is what you would call a dramatic game changer…
19 Sep 2016
A great programing interface (for R too): Jupyter notebooks
I recently started using jupyter notebooks for a project in which we are developing Python code for the computation of habitat connectivity measures (more on this project in the future).
in short, jupyter Notebooks are documents that combine text with live code, which are easy to share among collaborators.
Instalation of jupyter is relatively straightforward and well documented.
After having used it for python code, I recently discovered that it is also really powerful for use with R (i.e. the R in jupyteR).
I found this blog that convinced me to give it a go.
I hope to publish “soon” some results from our projects using the html export option from jupyter.
07 Aug 2016
The map below shows the different study areas for Renewable Reindeer, and is powered by Mymaps of Google:
19 Sep 2016
CLOSED
PhD on tourism and reindeer at the University of Stirling/NINA
A funded PhD project is available in the Conservation Science Group at the University of Stirling, UK, exploring the links between tourism and wild reindeer in Norway in the context of climate and land use change.
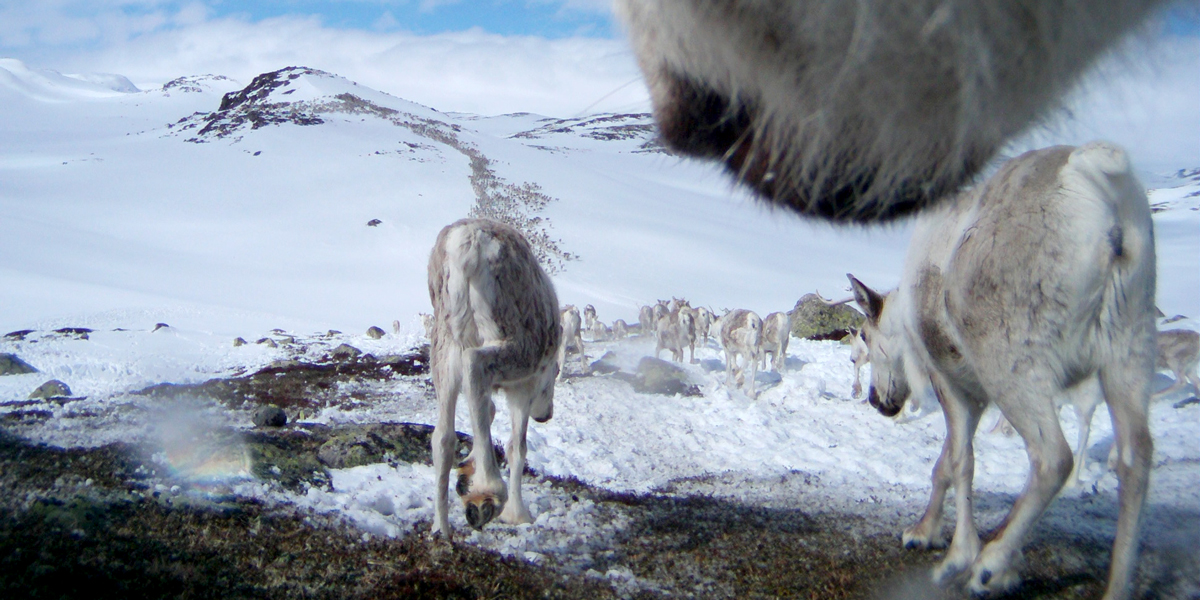

A funded PhD project is available in the Conservation Science Group at the University of Stirling, UK, exploring the links between tourism and wild reindeer in Norway in the context of climate and land use change. The project has a strong applied focus, aiming to inform on the sustainability of tourism and long‐term strategic planning for environmental management in Norway. The successful applicant will be supervised by staff based both at Stirling (Dr Nils Bunnefeld) and those based in the Norwegian Institute for Nature Research (Dr Bram Van Moorter, Dr Vegard Gundersen) and will work in close partnership with the Norwegian Wild Reindeer Center.
“Determining the impact of future tourism scenarios on sustainable use of mountains in the face of climate and land use changes for Norway’s emblematic wild reindeer.”
See full anouncement at the University of Stirling’s website.
19 Sep 2016

Journal of Animal Ecology, 85(1): 5-84.
Edited by Bram Van Moorter, Manuela Panzacchi, Francesca Cagnacci and Mark S. Boyce
Metabolism Special Feature Movements have long been identified as the glue linking individual behaviour to populations (Turchin 1998) and, ultimately, to species’ conservation and management. However, the field of movement ecology developed only recently, thanks to the rapidly increasing availability of both satellite tracking data of animal locations and of high-resolution environmental data layers. This led to a “perfect storm of opportunities” (Cagnacci et al. 2010) and to the rapid proliferation of a vast diversity of sophisticated analytical tools. However, for the classical schooled ecologists it has become increasingly challenging to find the most appropriate tool to answer a specific research question, and the current discourse in movement ecology is shifting in focus from the actual ecological questions to the analytical techniques to answer them. In conclusion, despite the great leap in the amount of information available, the way such information is used for ecology, management and conservation often falls short of its promises.
The motivation for this Special Feature is to refocus the discussions on key ecological questions in movement ecology and to provide guidelines for choosing the most appropriate analytical tools to answer them. The collection of 6 papers is the result of three years work starting from the summer of 2012, when we held a workshop in Evenstad, Norway, gathering lead scientists in movement ecology from Europe, America and Australia. The workshop was paralleled by a matching Summer School (International Research school for Applied Ecology, www.irsea.no), targeted at a broader audience. The Special Feature covers questions spanning from behavioral ecology to population dynamics, provides an overview of the state-of-the-art tools available to answer them, and proposes both theoretical and analytical advances to link individual movements to different kinds of spatially structured processes. In particular, GPS data are used to infer animal behavior (Guraire et al), migratory patterns (Cagnacci et al), identify the continuum between movement corridors and barriers (Panzacchi et al), quantify barrier permeability and proximity avoidance (Beyer et al), establish the link between individual movements, home range and habitat selection (Van Moorter et al), and scale up from habitat selection to population dynamics (Boyce et al) – as detailed in Borger et al. Online appendices allow the readers to access the codes and sample dataset needed to perform the analyses.
References
Turchin, P., 1998. Quantitative analysis of movement: measuring and modeling population redistribution in animals and plants. Sunderland: Sinauer Associates.
Cagnacci, F., Boitani, L., Powell, R.A. and Boyce, M.S., 2010. Animal ecology meets GPS-based radiotelemetry: a perfect storm of opportunities and challenges. Philosophical Transactions of the Royal Society B: Biological Sciences, 365(1550), pp.2157-2162.